Light sheet microscopy, such as SPIM (Selective Plane Illumination Microscopy) has been shown to be a valuable technology that performs especially well in samples that are too large for conventional microscopy, such as confocal. SPIM has spawned multiple variants, beam profiles and configurations. In simple terms for SPIM, optical sectioning is achieved by illuminating the sample with a sheet of light and generating fluorescence in a thin slice, which is then imaged with a fast and high resolution sCMOS camera. This allows for effective control of illumination of the specimen with a high signal to noise ratio. An aspect of SPIM and utilising sCMOS cameras is that these techniques generate approximately three orders of magnitude more data than a confocal microscope and therefore this can present a major burden of data transfer, storage and management.
In this article we look at how Jan Huisken and his research team (MPI, Dresden) took advantage of the light sheet microscopy approach of SPIM and the fast acquisition speed of the sCMOS camera to study the endoderm of the Zebrafish during early stages of development. Their research focusses on the coordination of cells in space and time that patterns the endorderm during the early stages of Zebrafish development. Early endoderm migration is of immense importance for organ formation, as defects in cell movement during gastrulation result in the duplication or mis-positioning of organs. Huisken and his team utilise compression of data and ‘on-the-fly’ processing to make this technique a success.
Multi-View Real-Time Imaging SPIM Configuration
Huisken and his team built a four-lens SPIM system (Fig 1) with integrated real-time image processing that delivers multiple views at high speed and is optimized for capturing the entire Zebrafish endoderm. The central unit of their setup consists of four identical objective lenses. The two detection lenses and their associated optics image the common focal plane onto two sCMOS cameras. A linear stage moves the sample along the detection axis through the focal plane, triggering the acquisition of the cameras every two microns. The two illumination lenses alternately illuminate the focal plane with laser light at every position during the scan. The cameras capture an image of the entire field of view ~1.2 mm at 62.5 fps.
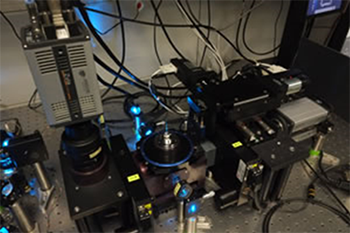

Figure 1. Four-lens SPIM setup and image acquisition.
Zebrafish embryos (5h post fertilization) were embedded in 1.5% low-melting agarose in a fluorinated ethylene propylene (FEP) tube. To enable visualization of the endoderm, the transgenic Zebrafish line Tg(sox17:EGFP), was used. The tube was mounted on the stage, dipping into the medium filled imaging chamber from the top. As the sample was moved through the light sheets, the cameras acquired the well resolved sectors of the embryo. This process took < 10s. Huisken found that by rotating the embryo by 45o at each time point to acquire a second, complementary view the quality of the image could be improved.
Managing the Data
The resultant high-resolution images of the entire endoderm were computed during the acquisition in real time. In their approach, a typical data set requires Gigabytes rather than Terabytes and offers visualization and evaluation of the cell migration in the context of entire embryos right out of the microscope. Taking insights from cartography they were capable of generating 2D map projections to visualize the entire embryo in a single image (Fig 2). In addition, real-time image registration eliminates any movement of the embryo in its chorion and registers all embryos to a common, predefined coordinate system.
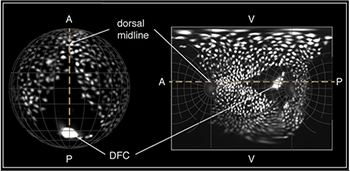
Figure 2. Spatial orientation of a Tg(Sox17:EGFP) embryo in 3D and on the final map projection.
This case study is based on a publication by Jan Huisken and his team and can be found at the following link: https://www.nature.com/articles/ncomms3207
Learn about the Dragonfly multimodal confocal and how it is suitable for developmental biology studies in the Dragonfly Whitepaper.
Andor Solutions for Light Sheet Microscopy
sCMOS cameras have proven to be ideal detectors for high-speed light sheet microscopy. Andor has a portfolio of sCMOS cameras that are highly suited to developmental biology studies:
The Andor Neo 5.5 and Zyla 4.2 PLUS series have been widely used since they offer the necessary large field of view and high resolution without compromising read noise or frame rate, and the 6.5 μm pixel size is ideally suited to light sheet microscopy. The new Sona back-illuminated sCMOS camera joins these models, with a similar sensor format, but with the higher QE of 95%.
The small pixel size of 6.5 micrometers of these cameras ensures sufficient oversampling of the pointspread-function, even for low magnification objectives that tend to be used for large specimens (e.g. 10x NA 0.3; 20x NA 0.5). The read noise is also very low meaning a low noise floor in the image. Zyla 5.5 can achieve a read noise down to 1.45 e– rms, while Zyla 4.2 has a read noise of 1.1 e- rms, both without signal amplification, while reading out 5.5 and 4.2 megapixels at 100 frames per second respectively. The latest Sona combines very low read noise down to 1.29 e- rms with higher QE of 95%, while offering 74 frames per second in full 16-bit operation.



