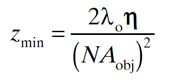...the application of deconvolution methods can always increase image quality, regardless of the source of the image...Deconvolve everything!
Mark B. Cannell, Angus McMorland, and Christian Soeller, Handbook of Biological Confocal Microscopy, Chapter 25
Deconvolution is a computational method of increasing the SNR (signal to noise ratio) and resolution of images captured on an imaging system. Its use predates the widespread use of confocal microscopy, but due to the lack of computing power at the time, was not commonly applied. Today’s computing power, particularly the massive parallelisation of GPUs (graphics processing units), has removed almost all barriers to entry, so that desktop PCs fitted with a suitable graphics card can execute deconvolution in almost real-time. The purpose of this technical note is to introduce the concept of deconvolution as a day-to-day imaging tool that should be routinely applied to any image captured on a microscopy system
Imaris 3D/4D image analysis software provides an optimal deconvolution module, ClearView-GPU, which is integrated into Imaris workflows and batchable. ClearView-GPU is executed directly from the Imaris software, equipped with crosssection preview mode, allowing instant feedback of the effects of a variety of deconvolution processing options and providing control over the result.
Key Features of Imaris ClearView-GPU™
- Powerful – GPU-accelerated enhanced Richardson-Lucy and Inverse filtering algorithms.
- Accurate - Accelerated Gibson-Lanni algorithm for accurate PSF estimates, supporting spherical aberration correction in deep specimens.
- Extremely fast - Optimized CUDA workflow for scorching performance.
- Quantitative - “Energy conservation” matches total photon content of raw data and results.
- Sharper – better optical sectioning and enhanced contrast even in deep specimens
- Integrated – Combine with acquisition for “real-time” feeling or operate on stored data.
- Innovative - Richardson-Lucy iteration acceleration with gradient-driven convergence in fewer cycles.
Image Formation
Image formation is the process by which an optical system projects an image of an object to a plane of observation or detection: the image is a spatial distribution of photons intended to represent the distribution of light emitted, reflected or transmitted from an object of interest. Image formation is described mathematically by the operation of “convolution”, in which the spatial distribution of light collected from the object is convolved with the instrument point spread function (PSF). PSF is considered a fundamental property of the physical imaging system and sets a limit to spatial resolution. Computer assisted imaging can exceed this physical limit: deconvolution processing is one such method. The PSF shape is limited by diffraction, usually at the pupil plane of the instrument. The narrower the PSF, the greater the spatial resolution and the larger the (numerical) aperture of the optical system. Convolution can be considered a mathematical description of blurring by the PSF, and it is this blurring we seek to “undo”’ in deconvolution.
In the case of fluorescence microscopy, the ability of a system to resolve an object is limited by the SNR and the resolution. Objects in the image may be affected in the following ways:
- If an object’s fluorescence intensity is too close to the noise floor of the detection system or background intensity of the sample, it will not be visible.
- If an object is smaller than the resolving power of the system, it will appear at least as large as the resolution limit of the system.
- If two objects are separated by less than the resolving power of the system, they will appear as one object.
Below we describe the deconvolution mathematically, and then visually, to support a more intuitive understanding. An important mathematical relationship relied upon in deconvolution is the Fourier Transform (FT). The FT is a way of describing distributions (commonly spatial or temporal), by a collection or set of alternative functions. To convert to the FT, we compute the amplitude and phase of a collection of spatial frequencies which describe the original function. The result is a set of pairs of sines and cosines at different spatial frequencies, each pair with an amplitude or constant of intensity. Calculation of this important function has been optimized over many generation and can now be computed extremely quickly with use of GPU power. It turns out that when distributions are expressed by their FT, that convolution of spatial distributions is represented by multiplication, while deconvolution is represented by division. That simplifies calculations quite a lot. Unfortunately, this simple relationship only holds true for the ideal noise-free case. For real imaging situations, noise makes this more challenging and then iterative techniques must be used.
Illustrating the simple case: the image received by the detector can be considered as made up from a collection of points of light, each of which has been convolved with the PSF. As a visual example, consider the following test pattern of objects (Fig.1):
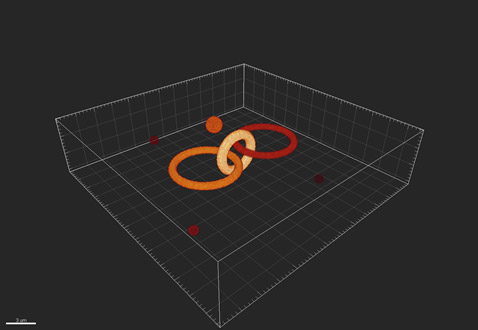

Figure 1 - An example of the true objects being imaged by the imaging system
After the light has passed through the optics of the system and received by the detector, it has been blurred or convolved so that the objects appear as shown in Fig. 2:
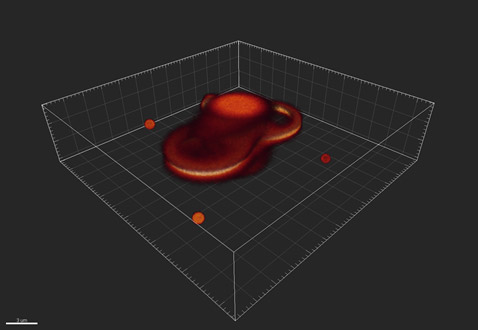

Figure 2 - An example of how the imaging system might distort, or convolve, the objects
The blurring function or point spread function is shown in Figures 3 and 4, which show a 3D view and 2D projection respectively. The rings in Figure 4 are known as Airy rings, and are characteristic of an imaging system with a circular aperture. In this case, 70% of the transmitted energy is contained in the central bright spot and the extent of this spot is known as one Airy unit, corresponding to 1.22*wavelength/NA. Where wavelength is the emitted wavelength and NA the limiting aperture of the optical imaging system.
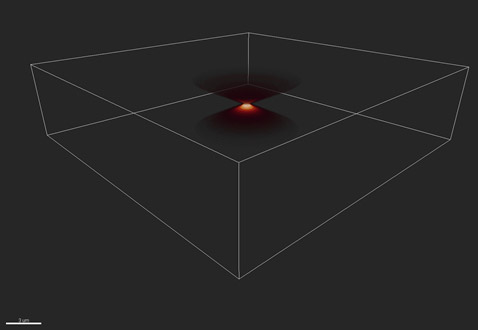

Figure 3 - The distortion of a single point, or Point Spread Function (PSF). Shown here in 3D

Figure 4 – The PSF in 2D shows a series of rings - known as the “Airy pattern” - with an “Airy disk” at its centre.
Since we know how the image is convolved with the PSF as it passes through the imaging system, we can apply the inverse of this process to deconvolve the image and recover both signal and resolution.
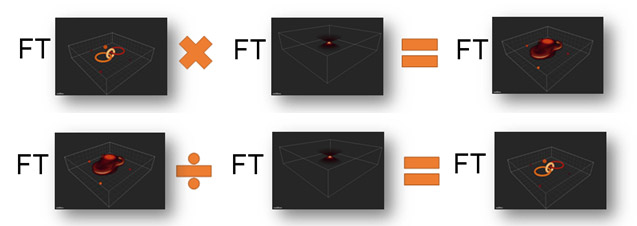
Figure 5 – Simplified explanation of image convolution and its inverse, deconvolution to recreate the original object. Note that FT represents the Fourier Transform, which allows us, in the noise-free case, to replace convolution with multiplication and deconvolution by division. Thus at top, we see the image formation process, in simplified form and below the image recover or deconvolution process.
Nyquist Sampling
In the early 20th Century, during the dawn of digital electronics, Harry Nyquist (and various others) realised that to accurately represent a continuous (or analogue) series in a discrete (or digital) way would require that the representation, or recording, of the continuous series be at least twice its frequency. An everyday example of this phenomenon is the recording of audio into digital formats at a frequency of at least 40 kHz (typically 44.1 kHz or greater is used). This is because the human auditory system (an analogue recording device) is generally not capable of detecting frequencies greater than 20 kHz. If the audio was recorded at less than 40 kHz, the high frequency treble sounds would be lost or worse, aliased into lower, spurious frequencies. The lower the recording frequency, the worse it would sound. We can represent this as follows
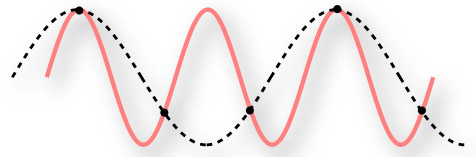
Figure 6 – Under sampling a continuous signal.
The red line is the continuous series we wish to record as a digital signal. The black dots are the frequency at which we record. The dashed line is thus the digital representation of the signal. It is obviously different than the original series!
A visual example of this is the moiré pattern seen in images that are resized or captured at low resolution:
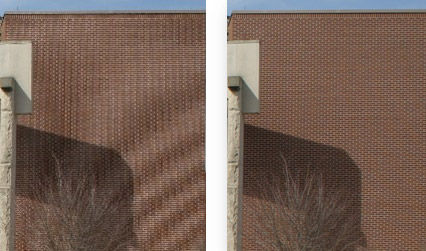

Figure 7 – The image on the left has not been captured at sufficient resolution and shows artefacts
In the case of imaging in three dimensions on a fluorescence microscope, we therefore need to set our image size (or more technically, our pixel size) and our step size (Z) appropriately.
The well-known Abbe equation tells us the limit of lateral (XY) resolution:
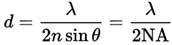
For example, using 525 nm light and a high NA objective, such as a 1.4 oil, this is 188 nm.
Nyquist sampling theorem tells us that our digital sampling should be at least twice this frequency (or half the distance), or that our pixel size (in the image plane) should therefore be no larger than 94 nm. If we are using a total magnification (C-mount and objective) of 100x, this equates to a 9.4 µm pixel on the camera.
The following formula tells us the limit of axial (Z) resolution:
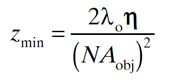
Where η is the refractive index of the medium.
Using the same example as before and 1.518 as the refractive index, this is 813 nm.
Nyquist sampling theorem tells us that our Z-step size should therefore be no larger than 406.5 nm, or 0.4 µm.
The take-home message is this: The more information that can be provided to the deconvolution processing, the better the resolution and SNR in the final image.
Within the usual restrictions of imaging with samples that may be sensitive to bleaching and/or phototoxicity, the smallest pixel size and smallest Z-step practical should be used. To get the best quality of your image and maximum deconvolution performance use the online calculators to assess the optimal voxel size.
ClearView-GPU™: Imaris Deconvolution Module
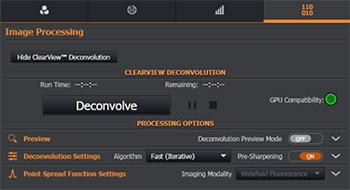
Figure 10 - Imaris Deconvolution module in the Image Processing Tab
Imaris ClearView-GPU™ module is found in the Image Processing section.
If a CUDA-compatible GPU and driver is found, you will see the information under Preferences\System and deconvolution processing will be up to fifty times faster than if it is executed on the CPU. Both options are supported in Imaris. Decenvolution should be selected from the drop down menu for every channel and preview window of adjustable size is available (volume projection in xy and yz and slice mode). Thanks to the preview mode users can get the instant feedback of the results and adjust the processing settings before applying them to an entire dataset.
It includes options for previewing a region of the image for instant feedback of the results, allowing the user to adjust the processing settings and see their effects before applying them to an entire dataset.

Figure 11 - Preview allows a small region to be deconvolved instantly
Algorithms and Point Spread Functions
Three processing algorithms are included with ClearView-GPU™ : Robust, Fast and Fastest. Each represents a balance between image quality and processing time.
- Robust is an iterative maximum likelihood estimator, will givinge the best results and is resistant to noise in the image.
- Fast is also an iterative method and uses the Van Cittert method to reduce the processing time, typically by a factor of two compared with Robust.
- Fastest is non-iterative and uses an inverse (Wiener) filter to further reduce the processing time, again typically by a factor of two compared with Fast.
In all cases, the dataset is split into chunks for processing on the GPU (defined by the GPU Processing Memory Limit setting in the Rendering menu of the Preferences area in Imaris). This means that there is no limit on the size of datasets that can be deconvolved, but that more GPU memory results in faster processing times.
ClearView-GPU™ includes five PSF models, all using a fast integrator to accelerate the Gibson-Lanni algorithm , used to estimate a robust 3D PSF, with aberrations. This is the best algorithm available today for PSF estimation.
- Spinning Disk Confocal
- Widefield Fluorescence
- Laser Scanning Confocal
- TIRF
Advanced Settings
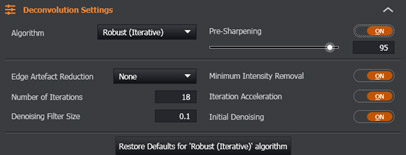
Figure 10 - Imaris Deconvolution advanced parameters module in the Image Processing Tab
The default settings have been carefully selected to ensure that the best results are given for most images. For users who wish to fine tune these settings, they can do so in the Deconvolution Settings section. It’s recommended to do this whilst in Preview Mode for an instant update of the effect of the setting.
Whilst beyond the scope of this document to provide a detailed explanation of all of them, Minimum Intensity Removal is pertinent in that users should disable this setting if they wish to maintain “energy conservation” and ensure that the photon (pixel) counts in their original and processed datasets are identical (< 1% discrepancy).
Results
| PSF measurements |
WF |
WF + decon |
DFly 40 |
DFly 40 + decon |
| Lateral FWHM (nm) |
245 |
185 |
238 |
139 |
| Lateral (XY) Projection |
 |
 |
 |
 |
| Axial (Z) Projection |
 |
 |
 |
 |
Table 1 - Comparison of imaging performance with the Dragonfly in widefield and confocal with 40 μm pinhole before and after deconvolution. Measurements were made with MetroloJ imageJ plugin for PSF analysis. 100 nm beads fluorescent were imaged at 488 nm laser excitation, with a Zyla 4.2 plus, 1X camera zoom and Nikon 60X/1.4 plan apo oil lens, Z step was 0.1 μm. FWHM is the full width at half maximum intensity across a line profile, a standard measure of the resolution.
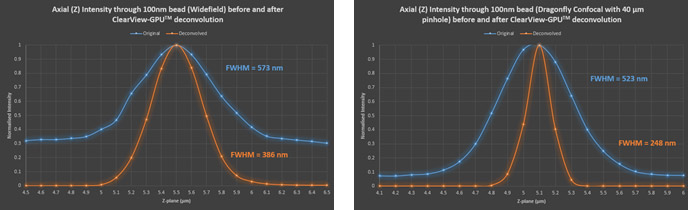
Figure 13 – Axial through-series (normalised) intensity profile of 100 nm bead showing the increase in SNR and decrease in FWHM (increase in resolution) after deconvolution in both widefield and confocal.
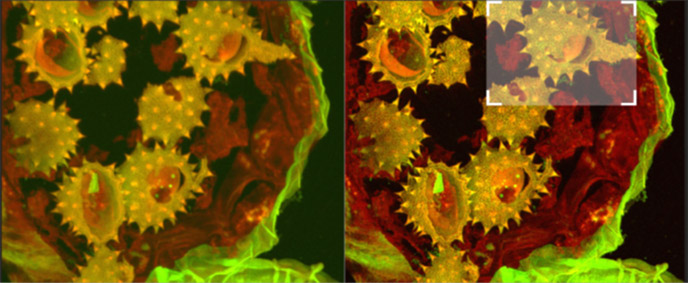
Fig. 14a - shows full field image of the daisy pollen grains before and after deconvolution, with merged channels viewed as a maximum intensity projection image. Note the contrast and detail enhancement in the deconvolved images. Because this is a bright robust specimen, we could extend exposure times to 250 ms, achieving high SNR to recover high spatial frequencies for deconvolution. Thus, we could exceed the Abbe diffraction limit in confocal imaging. But the density of the specimen is also a challenge to optical sectioning, as evidenced in the haziness of the single optical section in C. Daisy pollen maximum intensity projection of 488 and 561 channels before and after deconvolution with ClearView GPU™, imaged on Dragonfly with 488 and 561 laser excitation. A Zyla 4.2 PLUS was used with 1X camera zoom and Leica 100X/1.44 oil lens. Highlighted region shown below.
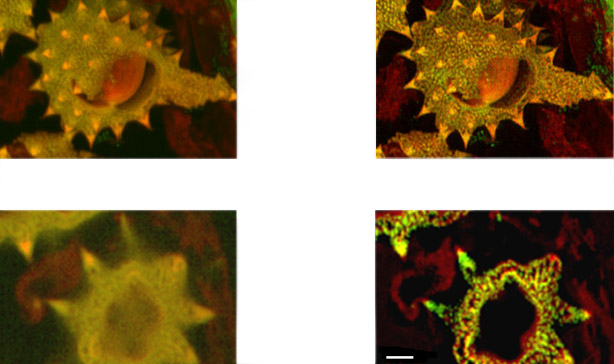
Fig. 14b (above) shows detail at full resolution from image in A above. Surface clarity is enhanced as well as sharpness and resolution improvement in individual pollen grain. Fig 14 c (below) shows single optical section from two channel daisy pollen Z series. ClearView-GPU enhances contrast, sharpens optical sectioning and provides clear channel separation in this thick bright specimen. Fine structures within the walls of the pollen grains become clearly visible. These punctate features are in the range 150-200 nm FWHM after deconvolution. Scale bar is 2 μm.
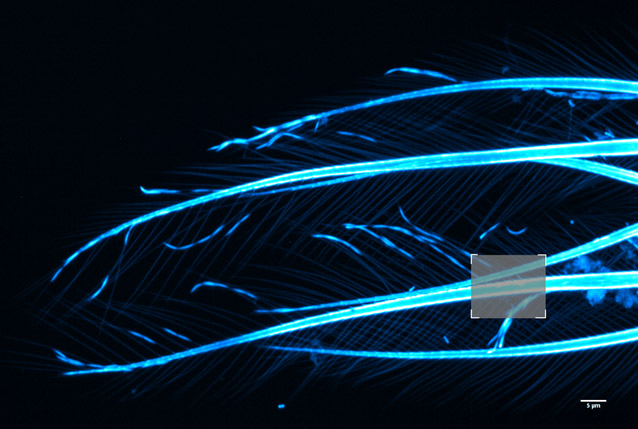

Figure 15 - Brine shrimp tail captured with a 40µm pinhole on Dragonfly. Objective is 63x 1.4. Camera is Zyla 4.2 Plus sCMOS with additional 1.5x magnification lens in Dragonfly. Highlighted region shown over in Fig. 16 before and after ClearView-GPU™ deconvolution.
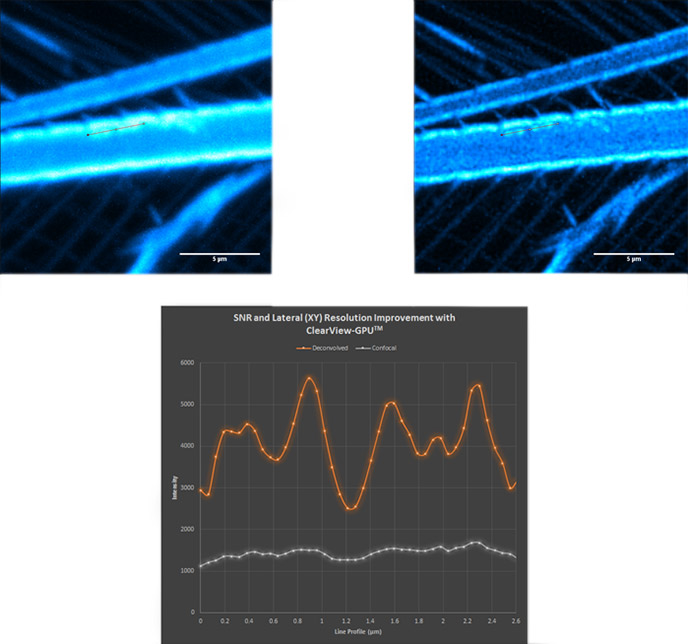
Figure 16 - Confocal (top left); deconvolved (top right) and line profile (bottom) showing increase in SNR and lateral resolution. Notice the haze along the line profile in the confocal image becomes clearly separated as four distinct objects after deconvolution.
Speed Comparison
Imaris ClearView-GPU™ has been engineered to ensure the highest speeds and functionality with the largest possible datasets.
Using identical hardware and as similar settings as possible, ClearView-GPU™ has been shown to be up to 50x faster than CPU-based methods and up to 10x faster than other leading GPU-accelerated packages, especially for larger datasets, even when iteration acceleration is disabled.
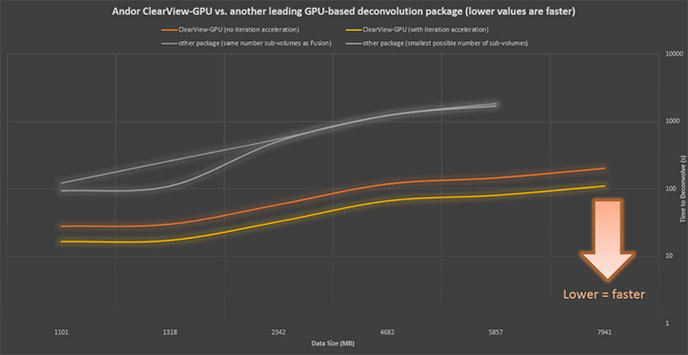
Figure 17 - Confocal (top left); deconvolved (top right) and line profile (bottom) showing increase in SNR and lateral resolution. Notice the haze along the line profile in the confocal image becomes clearly separated as four distinct objects after deconvolution.
Application Example
Besides the obvious visual benefit of deconvolution as described above, an additional more powerful benefit is how it can improve the ability to accurately analyse images using the wide range of tools available to researchers. To achieve the most accurate analysis, whether it be a simple point-to-point measurement, or the more sophisticated auto-detection of a subcellular structure and potentially tracking it over time, the most spatially conserved data (highest contrast and clearest boundaries) to the target elements are needed for investigation.
A good working example is that of vesicle tracking (fig. 18, below), such as in autophagy models for cancer research. The size of vesicles is in the range of a micron or less, and can be smaller than the pure resolution of an optical system without deconvolution. They can also be densely populated, as well as moving fast in a three-dimensional volume. The ability to discreetly detect individual vesicles and enable dedicated software analysis modules (e.g. Imaris TrackLineage) to autodetect and analyse them for parameters such as number, direction and distance travelled and speed of movement (figure 19), can be greatly enhanced by deconvolution.
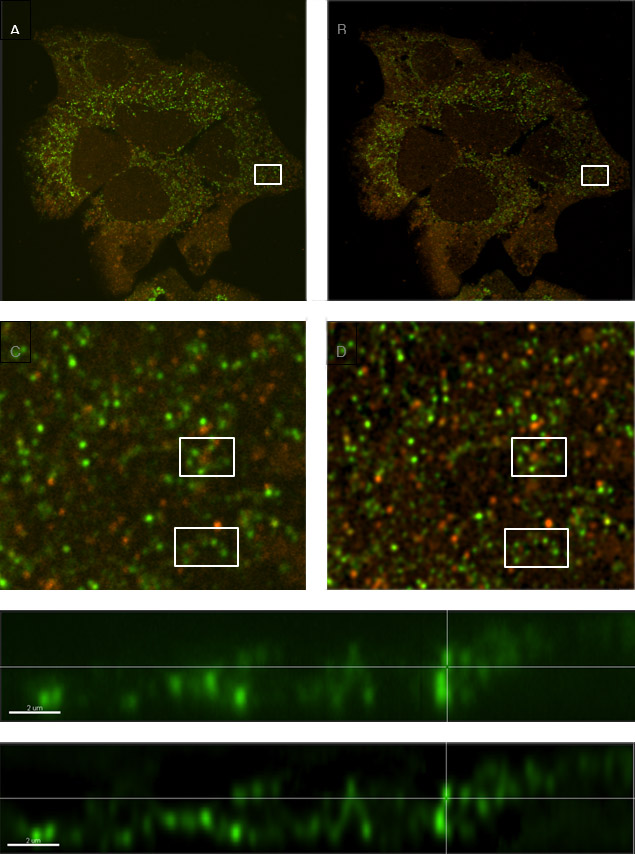
Figure 18 - Comparison of a sample with labelled vesicles imaged using Dragonfly confocal mode (A, C & E) and following ClearView-GPUTM deconvolution (B, D & F). C and D show zoomed regions of A and B respectively, and within those regions, clusters of vesicles are much clearer to see in the deconvolved image (D) compared to the raw confocal image (C). E and F are orthogonal (x,z) views demonstrating smaller diameter profiles with higher contrast in the deconvolved image (F) owing to the reassignment of light omitted from each vesicle back to its origin.
Discussion
We have shown that using ClearView-GPU™ deconvolution increases the resolution in all three dimensions, surpassing the Abbe limit in some cases, and increases the SNR for all imaging modalities. Features that were previously indistinguishable become clearly separated.
Deconvolution should be a standard first step preceding quantitative image analysis. Using Imaris ClearView-GPU deconvolution module is very time efficient because it overcomes the disadvantage of repeated file conversion, saving and opening the file in different software packages.
Another advantage of Imaris ClearView-GPU is that it can be combined with other image pre-processing filteres, viewed within one preview and executed in a batch mode for multiple images.
The original data is not modified as the deconvolved data is created in addition to, rather than replacing and can be used in quantitative studies if energy conservation is ensured using the advanced settings.
References
- Castleman, 1979; Agard and Sedat, 1983; Agard et al., 1989
- Image by Pluke - Own work, CC0, https://commons.wikimedia.org/w/index.php?curid=18423591
- CC BY-SA 3.0, https://commons.wikimedia.org/w/index.php?curid=644816
- James Pawley “Handbook of Biological Confocal Microscopy” 3rd Edition, Chapter 1, Springer, 2006
- S. F. Gibson, F. Lanni, “Experimental test of an analytical model of aberration in an oil-immersion objective lens used in three-dimensional light microscopy,“ J Opt Soc Am [A] 1991 Oct;8(10):1601-13.
- P.A. Jansson “Deconvolution of Images and Spectra”, 2nd edition, Dover Publications, Academic Press, 1997, ISBN-13: 978-0-486-45325-5